The Claesen group aims to functionally characterize molecular mechanisms that control bacterial interspecies and microbe-host interactions in the human microbiome. Bacteria use small molecule chemicals to mediate these interactions and the genetic information required for their production is typically encoded in one physical location of the bacterial chromosome, in biosynthetic gene clusters (BGCs). Using in silico techniques, we identified several widespread families of BGCs that we are now characterizing experimentally, prioritizing on the BGCs predicted to be involved in modulation of community composition or interaction with the host immune system. Our research will contribute to a better mechanistic understanding of the microbes that live in our gut and skin, leading to the discovery of druggable small molecules, new targets for antibacterial therapy and beneficial bacterial strains that can be employed for intervention therapies. The areas of expertise in the Claesen group include microbiology, bacterial genetics and synthetic biology, small molecule biosynthesis and biochemistry, and animal models for microbiome studies of bacterial infection, metabolic disease and cancer development.
Our group focuses on the following main themes:
Conversion of dietary flavonoids to bioactive monophenolic acids by the gut microbiota
Colorful fruits and vegetables are rich in plant metabolites called flavonoids. These compounds are attributed protective properties in cardiometabolic disease, diabetes and certain cancers. Since flavonoids themselves are poorly absorbed in systemic circulation, we are demonstrating that associated beneficial effects are actually caused by molecular transformations by certain members of the gut microbiota.

Discovery and engineering of Microbe-Microbe interactions
Bacteria produce specialized small molecules for interacting with other members of the community. We are mining the human gut and skin microbiomes for bioactive compounds that can be used for excluding pathogens or promotion of probiotic bacteria. We apply this knowledge to develop synthetic biology and engineering approaches for targeted community modulation.
Elucidating the biosynthesis and biological function of aryl polyenes
Aryl polyenes are specialized lipids that occur in the outer membranes of various Gram-negative bacteria. They confer a yellow pigmentation upon their producer strain and we showed their involvement in oxidative stress protection and biofilm formation. We’re currently investigating their biosynthesis and membrane attachment mechanisms as well as their role during host infection.

Claesen group publications:
31. Markley RL, I Johnston, V Bobba, S Ithychanda, G Mann, E Dester, K Ouyang, BP Matthew, and J Claesen (submitted). Oxidative stress induces E. coli aryl polyene expression, sensitizing the bacterial stress response and modulating the redox environment of innate immune cells. BioRXiv: 10.64898/2026.01.21.700819v1
30. Agudelo J, LJ Osborn, SD Mukherjee, A Franklin, A Adler, M Suryavanshi, B DeLucia, KB Schultz, W Massey, R Banerjee, A Horak, JK Nguyen, J Claesen, JM Brown, and AW Miller (submitted). Oxalate and dyslipidemia promote kidney stones and atherosclerosis dependent on gut microbiota.
29. Kumar M, R Son, SM Preston, R Glowacki, K Carr, J Kim, JZ Ma, PP Ahern, J Claesen, N Sangwan, F Rieder, and I Nemet (2026). The role of gut microbes in production of aromatic carboxaldehydes. Gut Microbes, 18(1): 2632979
28. Alqudah S, B DeLucia, LJ Osborn, RL Markley, V Bobba, SM Preston, T Thambidurai, L Hamidi Nia, CG Fulmer, N Sangwan, I Nemet, and J Claesen (2025). The diet-derived gut microbial metabolite 3-phenylpropionic acid reverses insulin resistance and obesity-associated metabolic dysfunction. Molecular Metabolism, doi.org/10.1016/j.molmet.2025.102272
27. Hamidi Nia L, S Alqudah, RL Markley, B DeLucia, V Bobba, J Elmallah, I Nemet, N Sangwan, and J Claesen (2025). The antimicrobial metabolite nisin Z reduces intestinal tumorigenesis and modulates the cecal microbiome in ApcMin/+ mice. Microbiol Spectr, 0:e01557-25, doi.org/10.1128/spectrum.01557-25
26. Elgharably H, J Claesen, N Sangwan, M Etiwy, P Houghtaling, GW Procop, NK Shrestha, B Griffin, JL Navia, LG Svensson, DJ Wozniak, and GB Pettersson (2025). In-vivo virulence of Staphylococcus aureus in native versus prosthetic left-sided valve endocarditis. JTCVS Open, doi.org/10.1016/j.xjon.2024.12.004
25. Alqudah S, and J Claesen (2024). Mechanisms of gut bacterial metabolism of dietary polyphenols into bioactive compounds. Gut Microbes, 16(1): 2426614
24. Claesen J, and JM Brown (2023). Predicting cardiovascular disease risk from gut microbial genes. mBio, e01970-23
23. Osborn LJ, K Schultz*, W Massey*, B DeLucia, I Choucair, V Varadharajan, R Banerjee, K Fung, AJ Horak, D Orabi, I Nemet, LE Nagy, Z Wang, DS Allende, BB Willard, N Sangwan, AM Hajjar, C McDonald, PP Ahern, SL Hazen, JM Brown#, and J Claesen# (2022). A gut microbial metabolite of dietary polyphenols reverses obesity-driven hepatic steatosis. Proc Natl Acad Sci USA, 119(48): e2202934119. (*equal contribution) (#co-corresponding)
22. Chambers LM, EL Esakov Rhoades, R Bharti, C Braley, S Tewari, L Trestan, Z Alali, D Bayik, JD Lathia, N Sangwan, P Bazeley, AS Joehlin-Price, Z Wang, S Dutta, M Dwidar, A Hajjar, PP Ahern, J Claesen, P Rose, R Vargas, JM Brown, C Michener, and O Reizes (2022). Disruption of the gut microbiota confers cisplatin resistance in epithelial ovarian cancer. Cancer Res, CAN-22-0455.
21. Hamidi Nia L, and J Claesen (2022). Engineered cancer targeting microbes and encapsulation devices for human gut microbiome applications. Biochemistry, doi.org/10.1021/acs.biochem.2c00251
20. Ozcam M, JH Oh, R Tocmo, D Acharya, S Zhang, S Ruiz-Ramírez, F Li, CC Cheng, E Vivas, FE Rey, J Claesen, T Bugni, J Walter, and JP van Pijkeren (2022). A secondary metabolite drives intraspecies antagonism in a gut symbiont that is inhibited by peptidoglycan acetylation. Cell Host Microbe, 30: 1-12.
19. DeLucia B*, S Samorezov*, MT Zangara, RL Markley, LJ Osborn, KB Schultz, C McDonald, and J Claesen (2022). A 3D printable device allowing fast and reproducible longitudinal preparation of mouse intestines. Anim Models Exp Med, 5: 189–196. (*equal contribution)
18. Osborn LJ, J Claesen#, and JM Brown# (2021). Microbial flavonoid metabolism: A cardiometabolic disease perspective. Annu Rev Nutr, 41: 433-454. (#co-corresponding)
17. Osborn LJ*, D Orabi*, M Goudzari, N Sangwan, R Banerjee, AL Brown, A Kadam, AD Gromovsky, P Linga, GAM Cresci, TD Mak, BB Willard, J Claesen#, and JM Brown# (2021). A single human-relevant fast food meal rapidly reorganizes metabolomic and transcriptomic signatures in a gut microbiota-dependent manner. Immunometabolism, 3(4): e210029. (*equal contribution) (#co-corresponding)
16. Johnston I, LJ Osborn*, RL Markley*, EA McManus, A Kadam, KB Schultz, N Nagajothi, PP Ahern, JM Brown, and J Claesen (2021). Identification of essential genes for Escherichia coli aryl polyene biosynthesis and function in biofilm formation. NPJ Biofilms Microbiomes, 7: 56. (*equal contribution)
15. Orabi D*, LJ Osborn*, K Fung, W Massey, AJ Horak III, F Aucejo, I Choucair, B DeLucia, Z Wang, J Claesen, and JM Brown (2021). A surgical method for continuous intraportal infusion of gut microbial metabolites in mice. JCI Insight, 6(9): e145607. (*equal contribution)
14. Claesen J, JB Spagnolo, S Flores Ramos, KL Kurita, AL Byrd, AA Aksenov, AV Melnik, WR Wong, S Wang, RD Hernandez, MS Donia, PC Dorrestein, HH Kong, JA Segre, RG Linington, MA Fischbach, and KP Lemon (2020). A Cutibacterium acnes antibiotic modulates human skin microbiota composition in hair follicles. Sci Transl Med, 12(570): eaay5445.
This paper was featured in the AAAS Podcast “How microbes compete for space on our face“.
13. Bell A, J Brunt, E Crost, L Vaux, R Nepravishta, CD Owen, D Latousakis, A Xiao, W Li, X Chen, MA Walsh, J Claesen, J Angulo, GH Thomas, and N Juge (2019). Elucidation of a sialic acid metabolism pathway in mucus-foraging Ruminococcus gnavus unravels mechanisms of bacterial adaptation to the gut. Nat Microbiol, 4(12): 2393-2404.
12. Ozcam M, R Tocmo, JH Oh, A Afrazi, JD Mezrich, S Roos, J Claesen, and JP van Pijkeren (2019). The gut symbionts Lactobacillus reuteri R2lc and 2010 encode a polyketide synthase cluster that activates the mammalian aryl-hydrocarbon receptor. Appl Environ Microbiol, 85(10): e01661-18.
11. Claesen J (2018). Topical antiseptics and the skin microbiota. J Investig Dermatol, 138(10):2106-2107.
Prior publications:
10. Ridaura V, N Bouladoux, J Claesen, E Chen, AL Byrd, M Constantinides, S Tamoutounour, MA Fischbach, and Y Belkaid (2018). Contextual control of skin immunity and inflammation by Corynebacterium. J Exp Med, 215(3): 785-799.
9. Smanski MJ, H Zhou, J Claesen, B Shen, MA Fischbach, and CA Voigt (2016). Synthetic biology to access and expand nature’s chemical diversity. Nat Rev Microbiol, 14(3): 135-149.
8. Thanapipatsiri A, J Claesen, JP Gomez-Escribano, M Bibb, and A Thamchaipenet (2015). A Streptomyces coelicolor host for the heterologous expression of Type III polyketide synthase genes. Microb Cell Fact, 14(1): 145.
7. Medema MH, R Kottmann, P Yilmaz, M Cummings, JB Biggins, K Blin, I de Bruijn, YH Chooi, J Claesen, RC Coates, et al. (2015). Minimum information about a biosynthetic gene cluster. Nat Chem Biol, 11(9): 625-631.
6. Claesen J, and MA Fischbach (2015). Synthetic microbes as drug delivery systems. ACS Synth Biol, 4(4): 358-364.
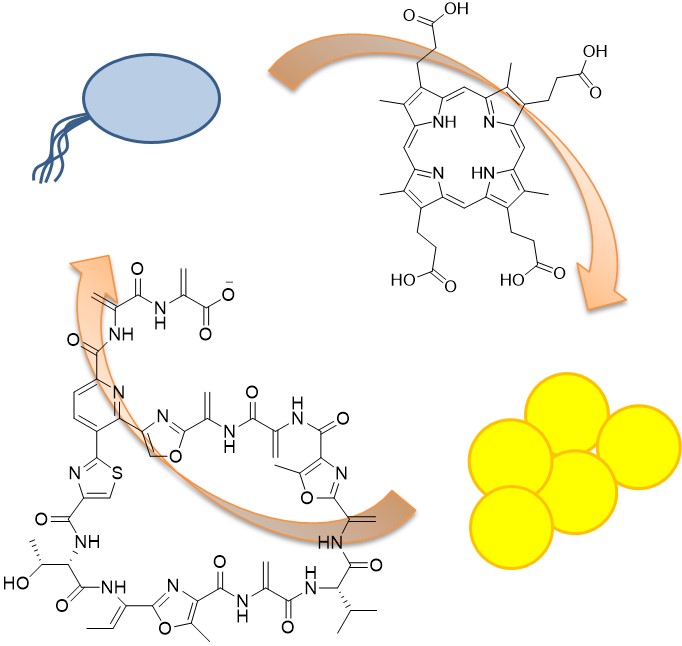
5. Wollenberg MS*, J Claesen*, IF Escapa, KL Aldridge, MA Fischbach, and KP Lemon (2014). Propionibacterium-produced coproporphyrin III induces Staphylococcus aureus aggregation and biofilm formation. mBio, 5(4): e01286-01214. (*equal contribution)
4. Cimermancic P*, MH Medema*, J Claesen*, K Kurita, LC Wieland Brown, K Mavrommatis, A Pati, PA Godfrey, M Koehrsen, J Clardy, BW Birren, E Takano, A Sali, RG Linington, and MA Fischbach (2014). Insights into secondary metabolism from a global analysis of prokaryotic biosynthetic gene clusters. Cell, 158(2): 412-421. (*equal contribution)
3. Claesen J, and MJ Bibb (2011). Biosynthesis and regulation of grisemycin, a new member of the linaridin family of ribosomally synthesized peptides produced by Streptomyces griseus IFO 13350. J Bacteriol, 193(10): 2510-2516.
2. Claesen J, and M Bibb (2010). Genome mining and genetic analysis of cypemycin biosynthesis reveal an unusual class of posttranslationally modified peptides. Proc Natl Acad Sci USA, 107(37): 16297-16302.
1. Goto Y, B Li, J Claesen, Y Shi, MJ Bibb, and WA van der Donk (2010). Discovery of unique lanthionine synthetases reveals new mechanistic and evolutionary insights. PLoS Biol, 8(3): e1000339.